The reg_lambda parameter in XGBoost controls the L2 regularization term, which helps prevent overfitting by adding a penalty to the model’s objective function based on the sum of squared weights.
By tuning reg_lambda, you can find the optimal balance between model complexity and generalization performance.
This example demonstrates how to tune the reg_lambda hyperparameter using grid search with cross-validation to find the best value that minimizes overfitting and improves the model’s ability to generalize to unseen data.
import xgboost as xgb
import numpy as np
from sklearn.datasets import make_regression
from sklearn.model_selection import GridSearchCV, KFold
from sklearn.metrics import mean_squared_error
# Create a synthetic dataset
X, y = make_regression(n_samples=1000, n_features=20, noise=0.1, random_state=42)
# Configure cross-validation
cv = KFold(n_splits=5, shuffle=True, random_state=42)
# Define hyperparameter grid
param_grid = {
'reg_lambda': [0, 0.1, 0.5, 1, 5, 10, 50, 100]
}
# Set up XGBoost regressor
model = xgb.XGBRegressor(n_estimators=100, learning_rate=0.1, random_state=42)
# Perform grid search
grid_search = GridSearchCV(estimator=model, param_grid=param_grid, cv=cv, scoring='neg_mean_squared_error', n_jobs=-1, verbose=1)
grid_search.fit(X, y)
# Get results
print(f"Best reg_lambda: {grid_search.best_params_['reg_lambda']}")
print(f"Best CV MSE: {-grid_search.best_score_:.4f}")
# Plot reg_lambda vs. MSE
import matplotlib.pyplot as plt
results = grid_search.cv_results_
plt.figure(figsize=(10, 6))
plt.semilogx(param_grid['reg_lambda'], -results['mean_test_score'], marker='o', linestyle='-', color='b')
plt.fill_between(param_grid['reg_lambda'], -results['mean_test_score'] - results['std_test_score'],
-results['mean_test_score'] + results['std_test_score'], alpha=0.1, color='b')
plt.title('Reg Lambda vs. MSE')
plt.xlabel('Reg Lambda (log scale)')
plt.ylabel('CV Average MSE')
plt.grid(True)
plt.show()
The resulting plot may look as follows:
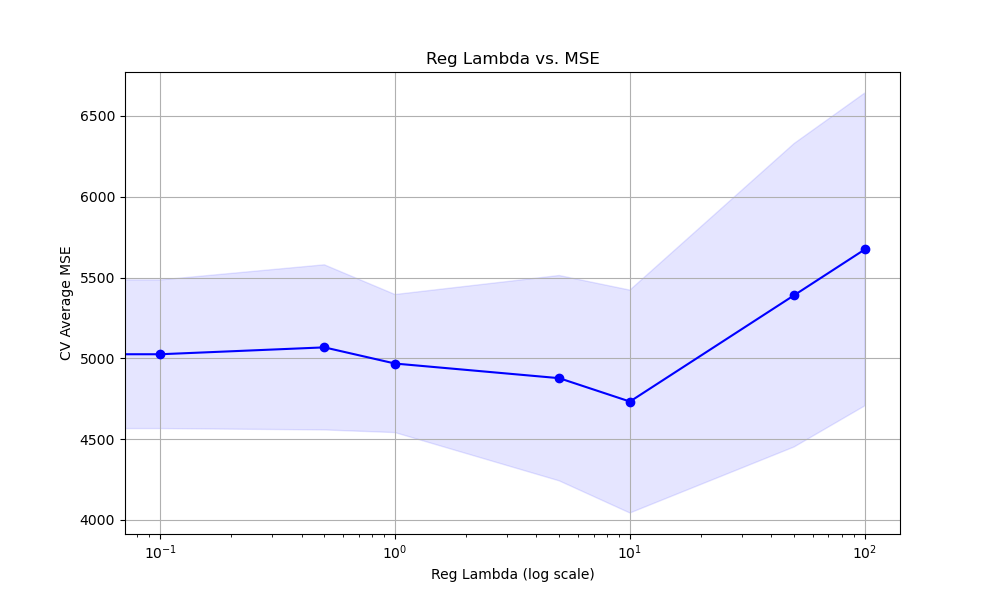
In this example, we create a synthetic regression dataset using scikit-learn’s make_regression function. We then set up a KFold cross-validation object to split the data into multiple folds for evaluation.
We define a hyperparameter grid param_grid that specifies the range of reg_lambda values we want to test. In this case, we consider values from 0 to 100, with a mix of small and large values to cover a wide range.
We create an instance of the XGBRegressor with some basic hyperparameters set, such as n_estimators and learning_rate. We then perform the grid search using GridSearchCV, providing the model, parameter grid, cross-validation object, scoring metric (negative mean squared error), and the number of CPU cores to use for parallel computation.
After fitting the grid search object with grid_search.fit(X, y), we can access the best reg_lambda value and the corresponding best cross-validation mean squared error using grid_search.best_params_ and grid_search.best_score_, respectively.
Finally, we plot the relationship between the reg_lambda values and the cross-validation average MSE scores using matplotlib. We retrieve the results from grid_search.cv_results_ and plot the mean MSE scores along with the standard deviation as error bars. We use a logarithmic scale for the x-axis to better visualize the range of reg_lambda values. This visualization helps us understand how the choice of reg_lambda affects the model’s performance and guides us in selecting an appropriate value.
By tuning the reg_lambda hyperparameter using grid search with cross-validation, we can find the optimal value that strikes a balance between model complexity and generalization performance. This helps prevent overfitting and ensures that the model performs well on unseen data.
